Our brains are what make us unique as humans, not just different from other animals but also from each other. The most recent studies tell us we have about 86 billion neurons, or brain cells, within our brain. The majority of the brain’s volume is taken up by axons, also known as nerve fibers.
These serve as long-distance connections or “communication highways” either within the brain or connecting the brain to the rest of our body. With so many highways, it should come as no surprise that the wiring of the brain is immensely complex and involves a lot of nerve fiber crossings to allow efficient connections.
Watch “Visualizing Crossing Fibers” on Labocine
Describe to us the imaging techniques used to make these brain renderings?
A medical imaging technique called Magnetic Resonance Imaging (MRI) allows us to take a detailed look into the body without disturbing the tissue or causing harm. The different magnetic properties of tissue types within our body allow us to depict the internal structure of the brain to about a one millimeter resolution.
With so many billions of cells within our brain, a resolution of one millimeter is still a very coarse picture. Diffusion MRI — the imaging technique used to obtain the data for these animations — is similarly limited in resolution, but the beauty of this technique is that it provides information on neuronal organization within each imaged voxel (a 3D variant of pixels), allowing a “virtual” approximation of the brain’s major wiring connections.
Apart from their aesthetic value, how do these animations help us understand broader questions about brain wiring and function?
The first scientific breakthrough in using diffusion MRI to get an idea of the brain’s wiring was about twenty years ago, and research into this field has since boomed with scientists ranging from mathematicians to neurosurgeons contributing to its ongoing improvement.
One of the biggest steps forward has been in extracting information on crossing nerve fibers from the MRI data, to best recover the complex wiring of the brain. How well we can do this is a strongly debated topic, and animations such as these help visualize and clarify what areas of the brain have such complex wiring.
Do you make animations as part of your research and where have they been shown?
After graduating from my Master’s degree at the VU University in Amsterdam in 2009, The Netherlands, I started a PhD research position in the University Medical Center in Utrecht, The Netherlands. Here I started working with diffusion MRI under the mentorship of Dr. Alexander Leemans, an incredibly talented person who got me hooked on this research field.
He developed software to analyze and visualize diffusion MRI data, called ExploreDTI, which is what all these animations are made in. Throughout my scientific career I have made many animations, initially starting just to see what was possible.
From there it gradually grew to showing footage to colleagues during meetings and as educational tools for student engagement. In 2013 the Dutch Brain Awareness week invited all Dutch neuroscientists to share their research images or videos with the public where my animation titled The Intergalactic Brain won the “Best Movie Award”.
I am currently employed at University College London, UK, where I work jointly in the Translational Imaging Group and Epilepsy department, where we use diffusion MRI in surgical planning on epilepsy patients. I use these types of animations to help familiarize clinicians with the do’s and don’t’s of the technique.
Why is film and imaging an important part of your scientific research?
The information we get from the diffusion MRI scans are almost as complex as the brains wiring itself. Our brain is very adept at interpreting visual cues, so showing the complex structure of the MRI data in a visual format helps us to understand our results. Also, animations are a much more effective way of reaching out to students and the public, which I hope will get other people interested in neuroscience and/or MRI.
SV: Most of what I use in these animations I have learned from my PhD project supervisor and mentor, Dr. Alexander Leemans.
The other developers on the ExploreDTI software tool, Ben Jeurissen in particular, have also made invaluable contributions without which I could not have made these animations. My colleagues at University College London help in discussing the best ways to convey the intended messages.
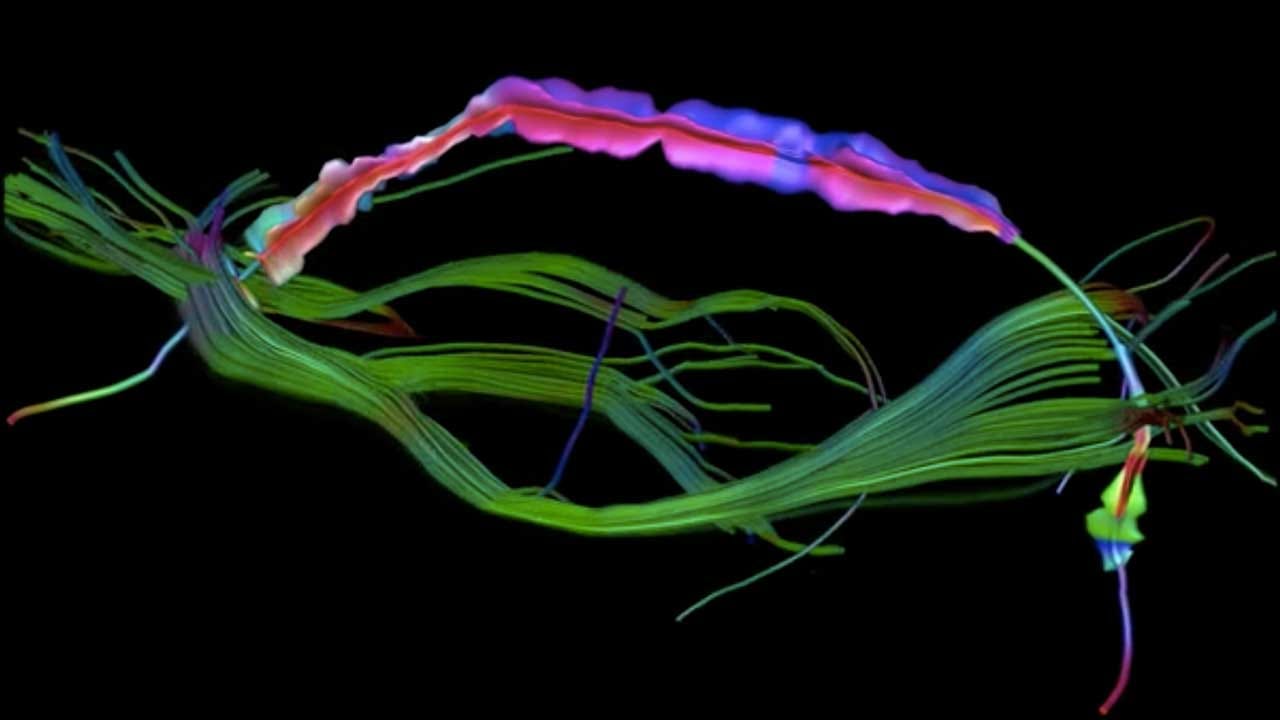
“Visualizing crossing brain fibers”
Diffusion MRI can provide information on the organization of neuronal tissue in vivo. Based on these images, fiber tract pathways can be reconstructed that represent the axonal wiring of the brain. Anatomical knowledge lets us select specific fiber bundles connecting two brain regions, in this case the arcuate fasciculus connecting two areas involved in language processing.
Visualization of all fiber tracts crossing such a fiber pathway can be informative to show the large-scale connectivity, but obscures the pathway of interest. By reducing these crossing fiber tracts to local representations, it becomes possible to visualize the complex tissue architecture locally along a fiber tract in a compact and elegant manner.
Copyright Sjoerd Vos, 2013. Created in ExploreDTI ©
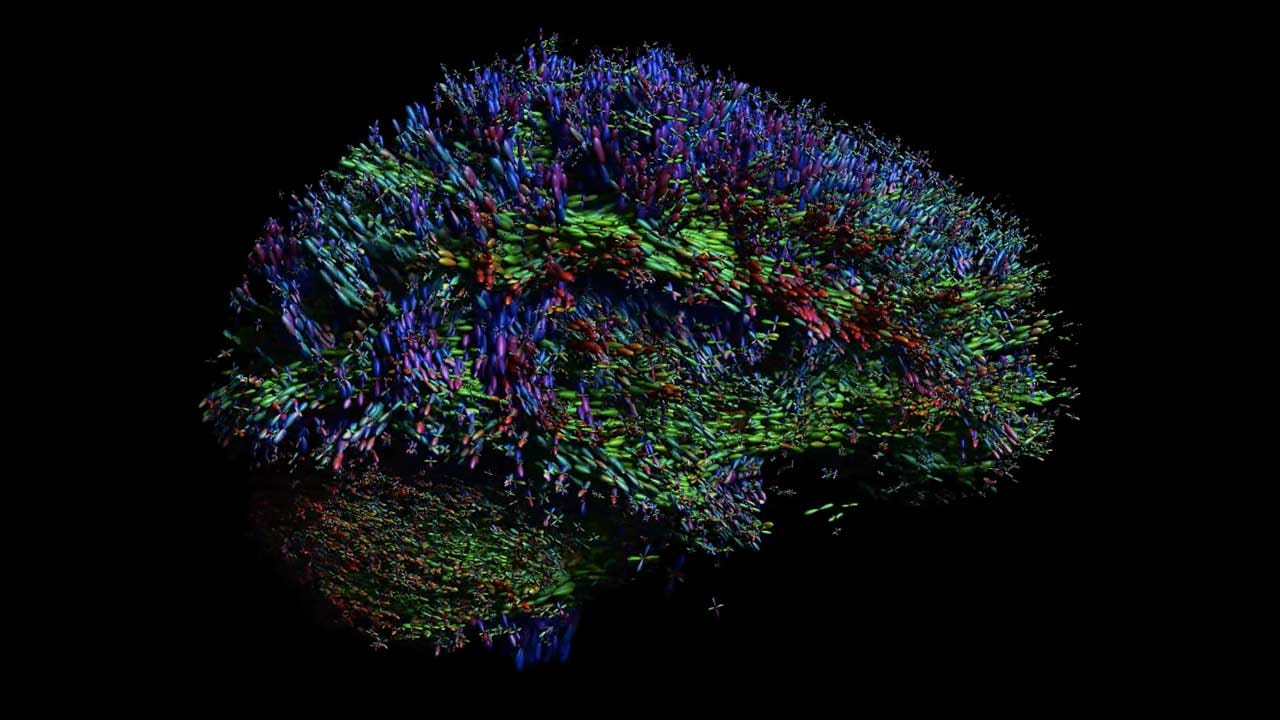
Watch “The Intergalactic Brain” on Labocine
With diffusion MRI, microstructural characteristics of white matter can be visualized at discrete positions throughout the brain. Resembling the scintillating objects in a galaxy, these multi-lobular star-shaped objects represent the local orientation of fiber tract pathways that interconnect distant brain regions.
This visualization — combined with our own brain power to ‘connect the dots’ — clearly shows the 3D architectural configuration of the brain.
Copyright ExploreDTI, 2013. Created with Alexander Leemans in ExploreDTI ©
“Brain wiring — simplicity or complexity”
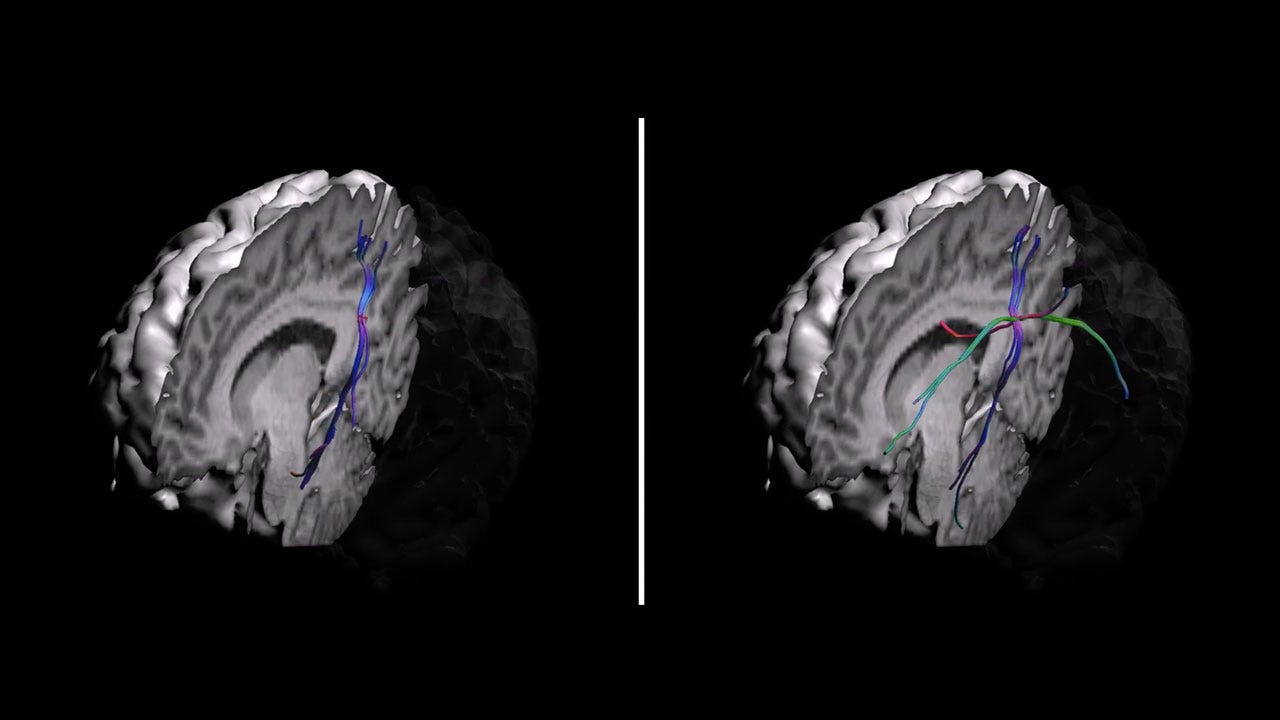
Watch Brain wiring — simplicity or complexity on Labocine
The scientific breakthrough in the 1990s that let us reconstruct the brain’s wiring did so simplifying the anatomy assuming that nerve fibers in each imaging voxel had a single main orientation.
With more advanced scanning options and processing options, it became possible to tease out crossing fibers: multiple nerve fibers in one voxel. The single voxel highlighted in this video is an example of where crossing fibers are present, and the reconstructed connections starting from that location are very different when using the initial method (left panel) or the crossing-fibers method (right panel).
Copyright Sjoerd Vos, 2016. Created using ExploreDTI ©
Sjoerd obtained his BSc in Medical Natural Sciences and MSc in Medical Physics from the VU University Amsterdam, the Netherlands. After graduation he started as a PhD student at the PROVIDI lab, part of the Image Sciences Institute in September 2009. The focus in his project was on the analysis of diffusion MRI data, more specifically diffusion tensor MRI (DT-MRI) and high angular resolution diffusion imaging (HARDI). These techniques can be used to investigate the microstructural organisation in the human brain. The aim of his project is to improve quantitative and qualitative analyses of these scans. During his PhD he spent six months at the Radiological Sciences Lab and the Center for Quantitative Neuroimaging at Stanford University, where he worked with Roland Bammer on high-resolution DWI acquisitions.
After his PhD, he did short-term post-doctoral research working closely with the Neurology department on improving the analysis of DTI data for Alzheimer’s disease patients. Parallel to that, he was also affiliated to the ENT department (ear, nose, throat) to improve the imaging and analysis of diffusion MRI data to image the auditory nerve.